Study: Molecular age of the eye determined for the first time
The researchers pointed out the eye can be a challenge to sample in living patients as, much like the brain, it is non-regenerative and obtaining a tissue biopsy would cause irreparable damage.
A team of researchers have mapped nearly 6000 proteins from different cell types within the eye by analyzing tiny drops of eye fluid that are routinely removed during surgery.1
According to the study, published this month in the journal Cell, the team used an AI model to create a “proteomic clock” from this data that can predict a healthy person’s age based on their protein profile.
Researchers noted the clock unveiled diseases such as diabetic retinopathy and uveitis that cause accelerated aging within specific cell types. Moreover, scientists also found proteins associated with Parkinson’s disease within eye fluid, which they say could offer a pathway to earlier Parkinson’s diagnoses.2
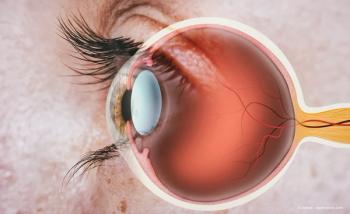
“What's amazing about the eye is we can look inside and see diseases happening in real time,” senior author Vinit Mahajan, a surgeon and professor of ophthalmology at Stanford University, said in the news release. “Our primary focus was to connect those anatomical changes to what’s happening at the molecular level inside the eyes of our patients.”
The researchers pointed out the eye can be a challenge to sample in living patients as, much like the brain, it is non-regenerative and obtaining a tissue biopsy would cause irreparable damage. As a result, an alternate method would require the use of liquid biopsies—samples of fluid taken from near the cells or tissues of interest.
According to the study, liquid biopsies can provide a snapshot of what proteins are situated in the area of interest. However, the researchers noted they have thus far been limited in their ability to measure large numbers of proteins within the small volumes of fluid, and they are also unable to provide information on which cells produced which proteins, which is important for diagnosing and treating diseases.1
In an effort to map protein production by different types of cells within the eye, Mahajan’s team used a high-resolution method to characterize proteins in 120 liquid biopsies taken from the aqueous or vitreous humor of patients undergoing eye surgery. The study noted that they were able to identify 5953 proteins—10 times the number of proteins previously characterized in similar studies. They created a a software tool dubbed TEMPO to trace each protein back to specific cell types.
The researchers also were examining the link between disease and molecular aging, the researchers built an AI machine learning model that can predict the molecular age of the eye based on a subset of 26 proteins. The model was able to accurately predict the age of healthy eyes but showed that diseases were associated with significant molecular aging.
For diabetic retinopathy, the degree of aging increased with disease progression and this aging was accelerated by as much as 30 years for individuals with severe (proliferative) diabetic retinopathy. These signs of aging were sometimes observable before the patient displayed clinical symptoms of the underlying disease and lingered in patients who had been successfully treated.1
Moreover, during the study the researchers uncovered proteins linked with Parkinson’s disease that are generally identified postmortem and current diagnostic methods aren’t capable of testing for them, which is a primary reason Parkinson’s diagnoses are so difficult. Screening for these markers in eye fluid could enable earlier diagnosis of Parkinson’s disease and later therapeutic monitoring.
The researchers pointed out in the study that data demonstrate aging could be organ- or even cell-specific, which could lead to improvements in precision medicine and clinical trial design.
“These findings demonstrate that our organs are aging at different rates,” first author and ophthalmologist Julian Wolf of Stanford University said in the news release. “The use of targeted anti-aging drugs could be the next step in preventative, precision medicine.”
Mahajan pointed out that if physicians are going to use molecular therapies, they should be characterizing the molecules in their patients.
‘I think reclassifying patients based on their molecular patterns and which cells are being affected can really improve clinical trials, drug selection, and drug outcomes,” Mahajan said.
The next step, according to the team, includes characterizing samples from a larger number of patients and a broader range of eye diseases. They also say that their method could be used to characterize other difficult-to-sample tissues. For example, liquid biopsies of cerebrospinal fluid could be used to study or diagnose the brain, synovial fluid could be used to study joints, and urine could be used to study the kidneys.1
The research was backed by Stanford University, the National Institutes of Health, Research to Prevent Blindness, the VitreoRetinal Surgery Foundation, the Lundbeck Foundation, and the BrightFocus Foundation.
Reference
1. Julian Wolfe, Ditte K. Rasmussen, Young Joo Sun, et al. Liquid-biopsy proteomics combined with AI identifies cellular drivers of eye aging and disease in vivo. Cell. Published October, 19, 2023; Accessed October 20, 2023. Doi: https://doi.org/10.1016/j.cell.2023.09.012
2. Molecular age of the eye determined for the first time. EurekAlert. Accessed October 20, 2023. Doi: https://www.eurekalert.org/news-releases/1004571
Newsletter
Keep your retina practice on the forefront—subscribe for expert analysis and emerging trends in retinal disease management.